Engineering Enzymes Without Crystal Structures: How AI and Structural Modeling Are Redefining Protein Design
AI and structural modeling are enabling enzyme engineering without crystal structures, unlocking faster, mechanism-aware, and data-driven protein design.
Introduction
Enzymes are central to modern biotechnology. From drug development and metabolic engineering to green chemistry and industrial catalysis, they enable highly specific and efficient biochemical transformations.
However, one of the most persistent challenges in enzyme engineering is the lack of high-resolution structural data.
A significant proportion of enzymes of interest do not have experimentally determined crystal structures, especially in ligand-bound or catalytically relevant conformations. This creates a fundamental bottleneck in rational enzyme design, where understanding active-site geometry, substrate binding, and catalytic mechanisms is essential.

Traditionally, this limitation has forced researchers to rely on indirect methods, approximations, or extensive experimental screening.
Today, advances in artificial intelligence and computational modeling are changing this paradigm.
The Structural Gap in Enzyme Engineering
Crystal structures have long been considered the gold standard for understanding enzyme function. They provide atomic-level insights into:
- active-site architecture
- substrate binding modes
- catalytic residue positioning
- conformational dynamics
However, obtaining crystal structures is not always feasible.
Many enzymes:
- are difficult to crystallize
- exist in multiple conformational states
- lack co-crystallized ligand complexes
- involve metal cofactors or transient intermediates
In particular, ligand-bound structures are often missing, which are critical for understanding how substrates orient within the active site and which reactive centers are involved in catalysis.
Without this information, enzyme engineering becomes significantly more complex.
The Traditional Workarounds and Their Limitations
In the absence of structural data, researchers have historically relied on:
1) Homology Modeling
Using known structures of related proteins as templates.
While useful, this approach depends heavily on sequence similarity and may fail when:
- homologs are distant
- active-site geometry differs significantly
- ligand interactions are not conserved
2) Directed Evolution
Iterative mutation and screening to identify improved variants.
Although powerful, this approach:
- is resource-intensive
- requires large experimental throughput
- provides limited mechanistic insight
3) Random Mutagenesis and Screening
Exploring sequence space without structural guidance.
This often results in:
- low hit rates
- unpredictable outcomes
- slow convergence toward optimal variants
These approaches, while foundational, are not well-suited for modern enzyme engineering challenges that require multi-objective optimization across activity, selectivity, stability, and safety.
A New Paradigm: Structure Without Crystallography
The emergence of AI-driven structure prediction and physics-based modeling has introduced a new paradigm.
It is now possible to generate high-confidence structural models of enzymes, even in the absence of experimental data.
AI-Based Structure Prediction
Recent advances in deep learning have enabled the prediction of protein structures with remarkable accuracy.
These models can:
- capture overall protein folds
- preserve active-site geometry
- provide a basis for downstream computational analysis
Importantly, they allow researchers to work with structural representations even when crystallographic data is unavailable.
Beyond Structure: Understanding Function Without Experimental Complexes
Predicting a protein structure is only the first step.
The real challenge lies in understanding how the enzyme interacts with its substrate.
This requires going beyond static models to capture:
- substrate binding orientation
- reactive center positioning
- catalytic residue interactions
- influence of cofactors such as metal ions
1) Docking and Binding Analysis
Computational docking methods can be used to generate plausible enzyme–substrate complexes.
However, standard docking approaches often fall short in enzyme systems, especially when:
- metal cofactors are involved
- active sites are flexible
- multiple binding modes are possible
2) Mechanism-Aware Modeling
To address these challenges, modern workflows integrate:
- active-site annotation
- catalytic residue mapping
- reactive center distance analysis
- pose comparison across multiple models
This allows researchers to move from simple binding predictions to mechanistic understanding of catalysis.
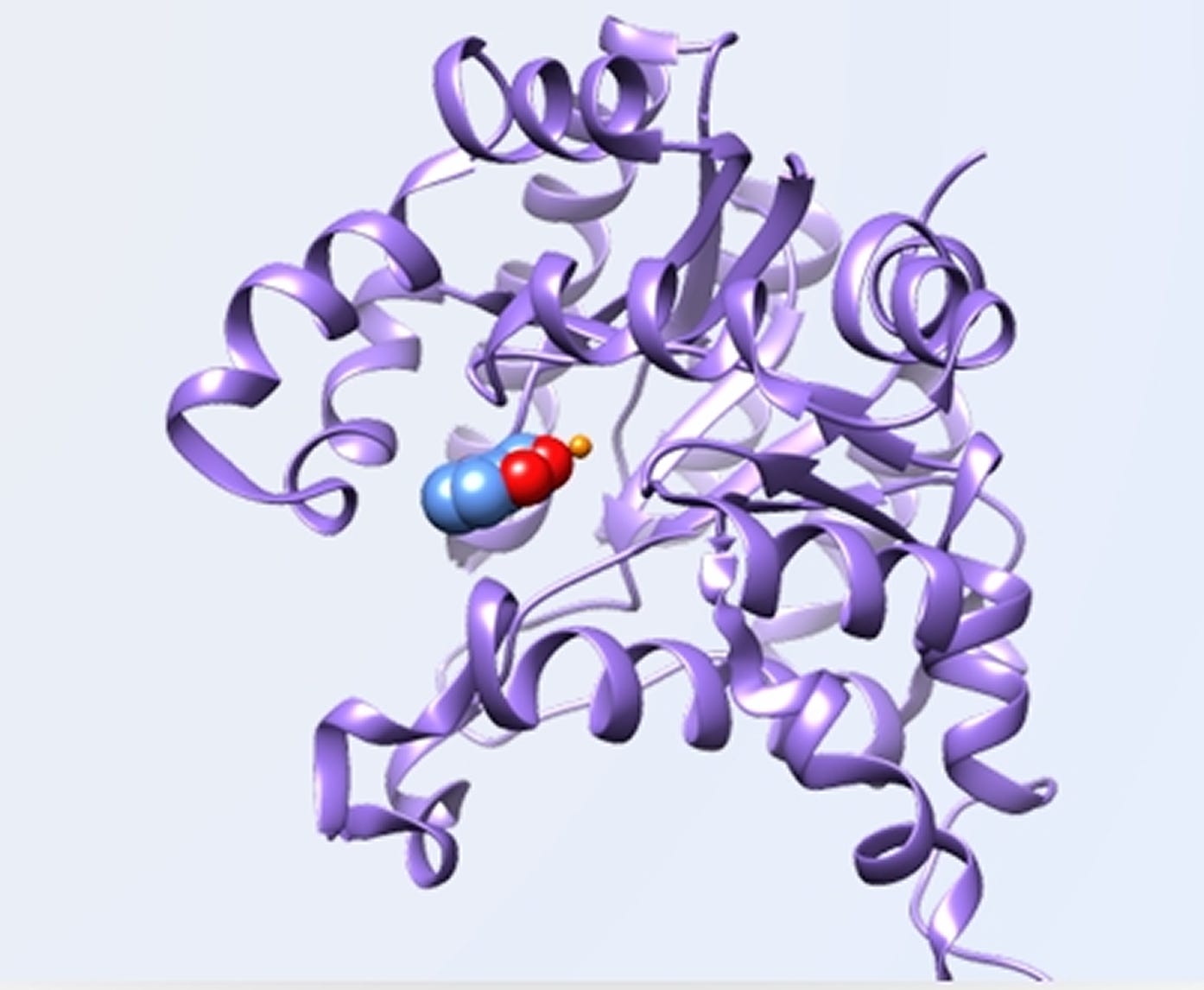
Case Study Insight: Engineering Regioselective Enzymes Without Crystal Structures
One of the most compelling applications of this approach is in the engineering of regioselective enzymes.
In many biocatalytic reactions, a substrate may contain multiple reactive sites. The enzyme’s ability to selectively act on one site over another determines the final product.
Without ligand-bound crystal structures, predicting regioselectivity is particularly challenging.
To overcome this, advanced computational pipelines can integrate:
- structure prediction for enzyme variants
- metal-aware docking to capture cofactor interactions
- mapping of reactive centers within the active site
- comparison of binding orientations across homologous enzymes
Such approaches enable the identification of enzyme variants that favor specific reaction pathways, even in the absence of experimental structural data.
The Role of AI in Navigating Structural Uncertainty
Artificial intelligence is not just enabling structure prediction. It is transforming how researchers explore enzyme design space.
1) Generative Design of Enzyme Variants
AI models can generate new enzyme sequences conditioned on:
- structural constraints
- substrate context
- catalytic requirements
This allows for the exploration of vast mutational landscapes that would be impossible to cover experimentally.
2) Multi-Parameter Optimization
Modern enzyme engineering requires balancing multiple factors simultaneously:
- catalytic efficiency
- substrate specificity
- structural stability
- solubility
- immunogenicity (for therapeutic enzymes)
AI-driven scoring frameworks can evaluate these parameters together, enabling more informed decision-making.
Medvolt’s Approach: Engineering Enzymes Without Structural Constraints
At Medvolt, enzyme engineering is approached as a structure- and mechanism-aware design problem, even in the absence of crystal structures.
Our platform integrates multiple layers of intelligence to overcome structural limitations:
1) Structure-First Modeling
High-fidelity 3D models are generated using advanced prediction and refinement workflows, preserving catalytic geometry and cofactor interactions where applicable.
2) Mechanism-Aware Analysis
We analyze:
- active-site architecture
- substrate binding orientation
- reactive center positioning
- catalytic residue engagement
This enables rational design decisions beyond simple sequence-level changes.
3) Metal-Aware Docking
For enzymes involving metal cofactors, such as Zn²⁺-dependent systems, we incorporate metal-aware docking protocols and reactive center mapping to accurately capture catalytic behavior.
4) Comparative Pose Analysis
Binding modes are evaluated across multiple homologs to identify conserved interaction patterns and prioritize functionally relevant conformations.
5) AI-Guided Variant Design
Generative models are used to design enzyme variants with desired properties, while multi-parameter scoring ensures that only high-confidence candidates are selected.
6) Physics-Based Validation
Top candidates undergo molecular simulations to validate stability, binding behavior, and dynamic interactions before experimental testing.
Together, this forms a closed-loop AI–experimental workflow that reduces uncertainty and accelerates enzyme discovery.
Implications for Biotechnology and Drug Discovery
The ability to engineer enzymes without crystal structures has far-reaching implications.
1) Accelerated Discovery
Researchers can move from concept to candidate faster by eliminating reliance on experimental structure determination.
2) Expanded Design Space
Enzyme families that were previously inaccessible due to lack of structural data can now be explored.
3) Improved Selectivity and Efficiency
Mechanism-aware modeling enables precise control over reaction outcomes, including regioselectivity and substrate specificity.
4) Reduced Experimental Burden
By prioritizing high-confidence candidates computationally, the number of required wet-lab experiments can be significantly reduced.
The Future: From Data Gaps to Design Opportunities
The absence of crystal structures is no longer a fundamental barrier to enzyme engineering.
Instead, it is becoming an opportunity to leverage advanced computational tools for more efficient and informed design.
As AI models continue to improve and integrate with physics-based simulations and experimental feedback, enzyme engineering will become increasingly predictive, scalable, and accessible.
The field is moving away from trial-and-error approaches toward rational, data-driven design frameworks.
Conclusion
Engineering enzymes without crystal structures was once considered a major limitation.
Today, it is a challenge that can be systematically addressed through the integration of AI, structural modeling, and mechanistic analysis.
By combining predictive modeling with experimental validation, it is now possible to design enzymes with high precision, even in the absence of traditional structural data.
At Medvolt, this philosophy underpins our approach to enzyme engineering, enabling the design of high-performance enzymes for therapeutic, industrial, and biocatalytic applications.
The future of enzyme engineering is not constrained by missing structures.
It is defined by how effectively we can model, understand, and design around them.