Unveiling SEE-CITE: Precision Mapping of Drug-Protein Interactions with an AI-Driven Paradigm
SEE-CITE introduces an AI-driven approach to map drug-protein interactions with high precision, enabling faster and more accurate drug discovery workflows.
Introduction: Charting New Territories in Drug-Protein Interaction Mapping
Drug discovery has always hinged on understanding how molecules interact within the complex signaling networks of living systems. At the heart of this challenge lies the intricate process of mapping drug-protein interactions (DPIs). These interactions dictate therapeutic efficacy, off-target effects, and the fundamental pharmacokinetics of a compound. While traditional experimental approaches like X-ray crystallography or nuclear magnetic resonance (NMR) spectroscopy provide high-resolution insights into these interactions, they often fall short in scalability, cost-efficiency, and precision when applied to high-throughput screening.
Enter SEE-CITE (Selective Energy Enrichment-Chemical Interaction Targeting and Enumeration), a cutting-edge technology designed to map DPIs with unsurpassed precision. Combining the computational might of artificial intelligence (AI) with advanced molecular topology methods, SEE-CITE offers an integrative platform that refines both target identification and validation workflows. By enriching data with precision-driven algorithms, SEE-CITE is helping researchers link the molecular footprint of small molecules with functional therapeutic outcomes.
The Need for Targeted Precision in Drug Discovery
The drug discovery landscape faces multifaceted bottlenecks. On average, it takes more than a decade to bring a drug from initial concept to market approval, at a cost often exceeding $2 billion. One of the most formidable challenges lies in pinpointing DPI at the earliest stages of development to avoid costly late-stage failures due to toxicity or lack of efficacy.
Traditionally, brute-force experimental techniques have been used to identify binding pockets on proteins or assess binding affinities of ligands to targets. However, such methods often produce incomplete datasets that either miss transient binding sites or fail to accurately capture conformational protein dynamics. Computational chemistry tools, such as molecular docking and free energy perturbation (FEP) simulations, have bolstered predictions, but their reliance on static snapshots often limits their practical applications.
SEE-CITE bridges these gaps by integrating real-time experimental data with predictive AI models to dynamically map binding sites and detect subtle interaction signatures. This enhanced precision promises to transform the way pharmaceutical and biotech firms prioritize and optimize lead candidates.
How SEE-CITE Works: Breaking Down the Technology
SEE-CITE operates at the convergence of several cutting-edge technologies:
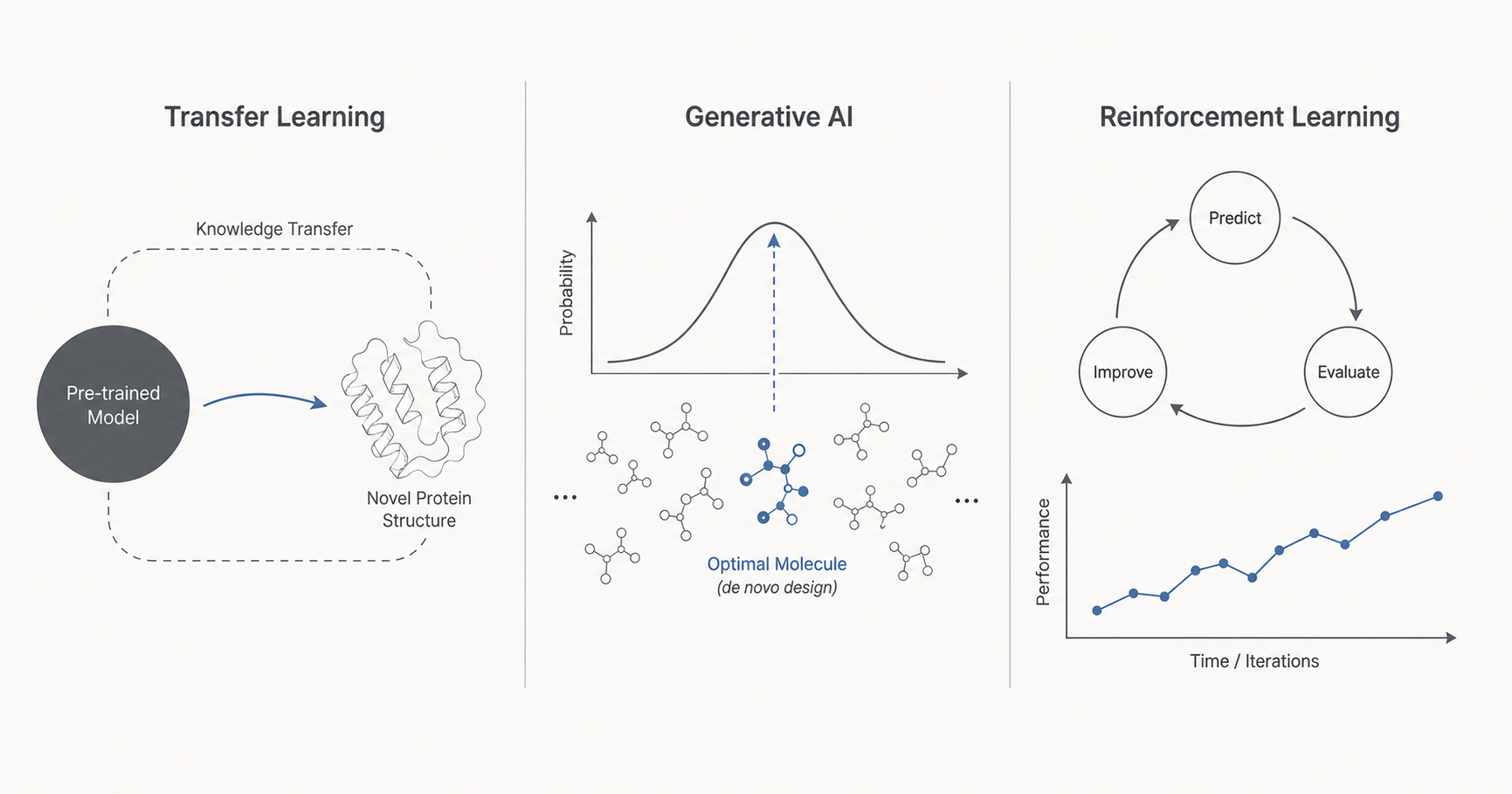
1. Selective Energy Enrichment for Binding Site Detection
This initial step in SEE-CITE employs selective energy enrichment algorithms, which identify thermodynamically favorable binding regions on proteins. Unlike conventional docking methods that focus narrowly on static crystal structures, SEE-CITE incorporates protein flexibility into its energy profile modeling, ensuring accurate consideration of both primary and allosteric binding sites.
2. Chemical Interaction Targeting with AI-Augmented Precision
Beyond binding energy screening, SEE-CITE leverages deep learning neural networks to predict and classify interaction motifs. By analyzing large multi-omics datasets, the AI algorithms can identify conserved motifs that consistently appear in protein-ligand interactions across diverse biological systems. This data-driven approach circumvents biases introduced from using isolated structural databases.
3. Interaction Enumeration for Holistic Mapping
SEE-CITE’s advanced enumeration capabilities enable researchers to visualize not just primary protein-ligand interfaces but also their cascading effects on downstream signaling. By overlaying interaction data onto cheminformatics pathways, researchers gain a full-spectrum view of DPI and its systems-level implications.
Applications and Advantages of SEE-CITE
The implications of SEE-CITE are vast, influencing multiple domains of life sciences, from drug discovery to enzyme engineering and beyond. Below are some specific ways that SEE-CITE is driving innovation:
1. Accelerating Hit-to-Lead Optimization
Typically, the transition from hit identification to lead optimization involves iterative cycles of synthesis and testing. SEE-CITE’s ability to rapidly assess the binding affinity and specificity of multiple compounds simultaneously reduces the need for costly trial-and-error steps. Researchers can focus on compounds that display the best therapeutic profiles in silico, dramatically shortening project timelines.
2. Unraveling Allosteric and Transient Targets
Transient or allosteric binding sites are challenging to identify due to their dynamic nature. SEE-CITE’s incorporation of protein dynamics into its modeling pipeline allows it to uncover such cryptic sites, paving the way for novel therapeutic interventions in areas such as oncology, neurodegenerative disorders, and immunology.
3. Enhancing Polypharmacology Development
Diseases with complex etiology often require drugs capable of modulating multiple targets simultaneously. SEE-CITE’s enumeration capabilities enable the design of polypharmacological agents by mapping potential on-target and off-target interactions in a systems biology context, minimizing adverse outcomes while maximizing therapeutic breadth.
4. Enabling Rational Enzyme Engineering and Biocatalysis
Beyond pharmaceutical applications, SEE-CITE plays a vital role in enzyme engineering. By elucidating enzyme-substrate binding sites with atomic-level accuracy, this platform enables the rational design of biocatalysts optimized for specific industrial, environmental, or pharmaceutical applications.
The Role of AI in Enhancing SEE-CITE’s Impact
At the core of SEE-CITE lies the transformative potential of AI. By integrating training datasets drawn from multi-omics platforms (e.g., genomics, proteomics, and metabolomics), AI algorithms provide unparalleled context for analyzing drug-protein interaction networks. Machine learning enhances the ability to define complex biochemical relationships that are often missed by humans or traditional computational tools.
In particular:
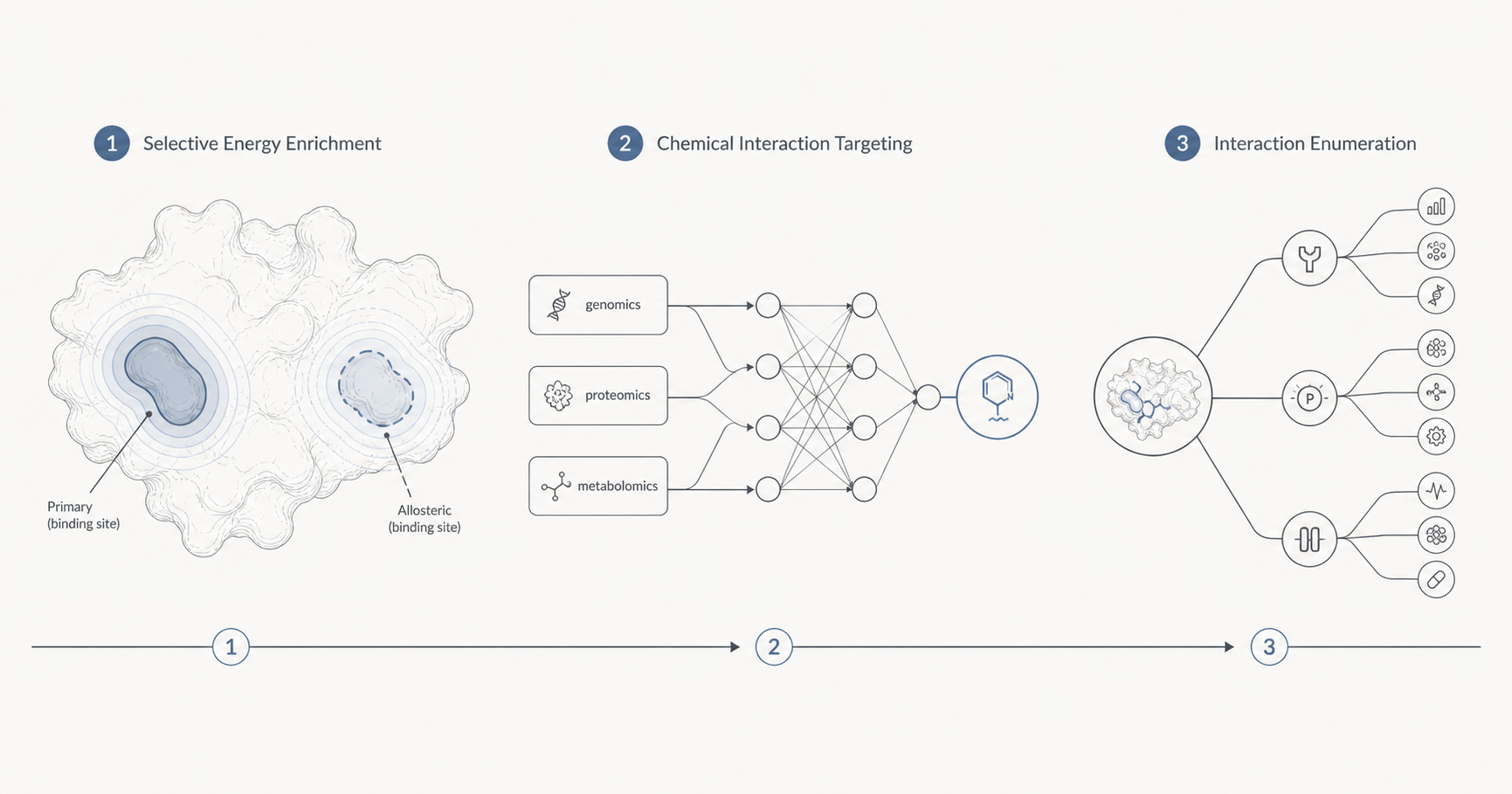
- Transfer Learning enables the application of existing models to novel proteins, reducing the need for extensive data curation.
- Generative AI Models support the de novo design of small molecules with optimized binding profiles by using probabilistic approaches rather than deterministic ones.
- Reinforcement Learning optimizes drug design pipelines by continuously improving predictions based on iterative feedback loops.
Future Directions and Challenges
While SEE-CITE holds immense promise, it is not without challenges. The accuracy of its prediction models ultimately depends on the quality, diversity, and size of datasets used to train its algorithms. Developing methods for cleaning and normalizing enormous datasets is an ongoing effort among computational chemists.
Moreover, as the complexity of therapeutic targets increases (e.g., intrinsically disordered proteins or membrane proteins), SEE-CITE’s models will require further refinement to maintain performance. Collaborations between AI leaders and experimental researchers will be essential to ensure a robust feedback loop underpins this iterative process.
Conclusion: SEE-CITE as a Paradigm Shift
SEE-CITE offers more than just incremental progress—it redefines how researchers approach drug-protein interactions. By marrying AI-driven computational tools with experimental insights, the technology equips scientists, pharmaceutical enterprises, and biotech innovators with an unprecedented level of precision in R&D workflows. Whether applied to therapeutic design, enzyme engineering, or molecular systems biology, SEE-CITE is poised to be a cornerstone for the next era in life sciences innovation.
Through partnerships with forward-thinking tech companies like Medvolt, the promise of targeted precision in drug discovery is becoming an attainable reality. With tools like SEE-CITE, tomorrow’s breakthroughs are within reach today, transforming disease treatment, prevention, and ultimately, human health.