Why Most AI Drug Discovery Pipelines Fail
AI drug discovery pipelines fail not due to weak models, but due to fragmentation, poor integration, and lack of physics-based validation.
Understanding the Gap Between Promise and Reality in Computational Pharma
Artificial intelligence has become one of the most widely discussed forces in modern drug discovery. Over the past decade, AI has been positioned as a transformative technology capable of reducing timelines, lowering costs, and improving success rates across the pharmaceutical pipeline.
From target identification to molecule design and optimization, AI models are now being applied across nearly every stage of discovery. The expectation is clear: smarter algorithms should lead to better drugs, faster.
Yet despite this surge in adoption, a difficult question remains. If AI is so powerful, why do most AI-driven drug discovery pipelines still fail to deliver consistent, real-world outcomes?
The answer lies not in the limitations of AI itself, but in how it is being used.
The Promise of AI in Drug Discovery
The appeal of AI in drug discovery is rooted in its ability to process complex, high-dimensional data and uncover patterns that are difficult for humans to detect.
Modern AI systems can:
- predict protein structures with high accuracy
- generate novel molecular scaffolds
- identify disease-associated biological targets
- optimize chemical properties across multiple parameters
These capabilities have enabled a new generation of computational approaches that promise to accelerate discovery and reduce reliance on trial-and-error experimentation.
However, these models often operate under an implicit assumption: that accurate predictions at individual stages will translate into successful outcomes across the entire pipeline.
In practice, this assumption frequently breaks down.
The Fragmentation Problem
One of the most common reasons AI drug discovery pipelines fail is fragmentation.
In many organizations, AI models are developed and deployed as independent tools. A target identification model operates separately from a molecule generation system, which in turn is disconnected from docking, simulation, and optimization workflows.
This creates a pipeline that is technically advanced, but scientifically disjointed.
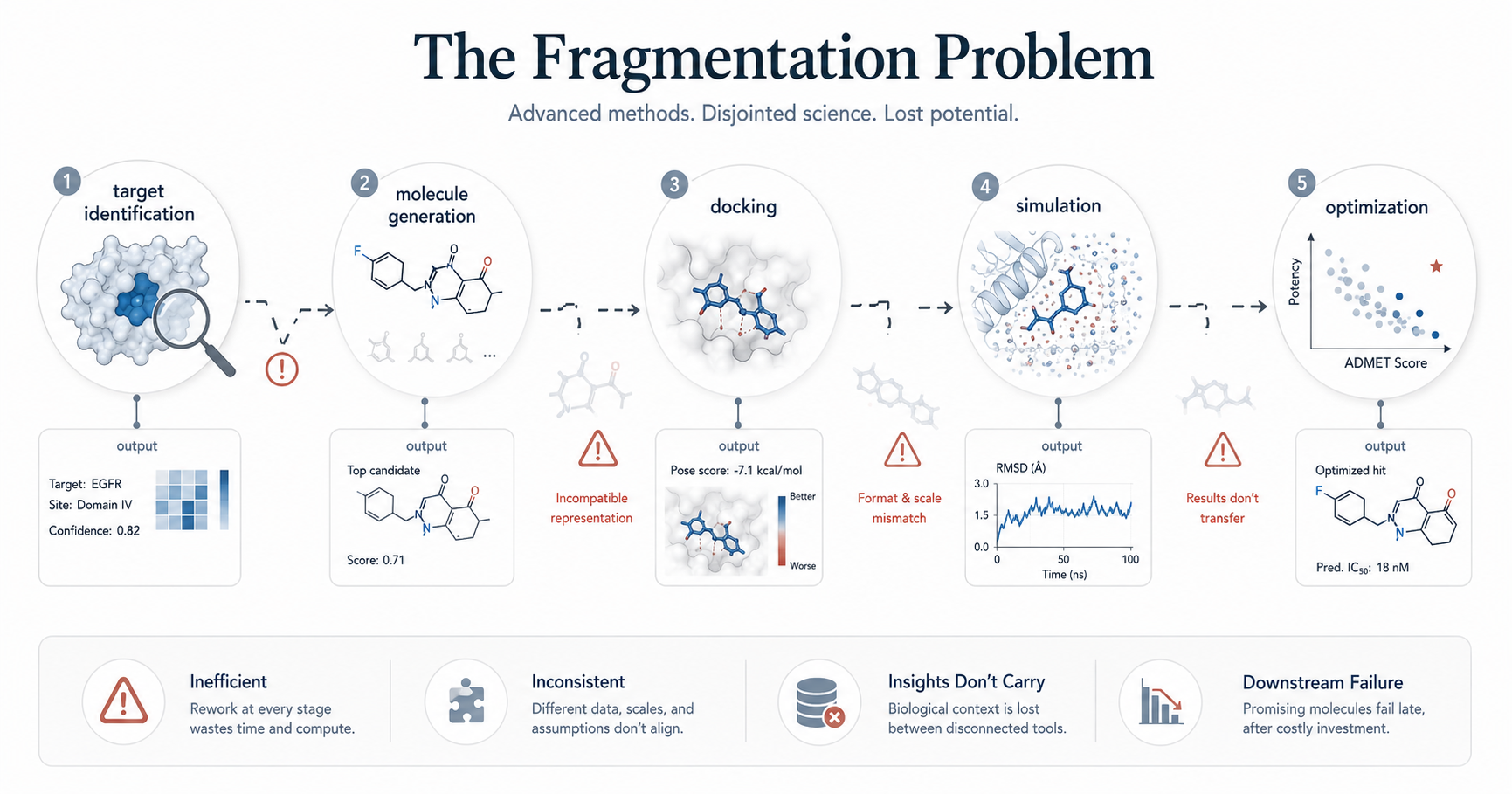
A molecule generated by one model may not be compatible with the assumptions of another. Biological insights derived from one dataset may not carry through to downstream stages. Each model optimizes for its own objective, without a unified understanding of the overall discovery goal.
As a result, promising predictions at one stage often fail when evaluated in a broader context. This fragmentation introduces inefficiencies, inconsistencies, and ultimately, failure.
The Overreliance on Single Metrics
Another major limitation in AI pipelines is the tendency to optimize for narrow objectives.
For example, a model may be trained to maximize predicted binding affinity or generate molecules with favorable docking scores. While these metrics are useful, they represent only a small part of what determines a successful drug candidate.
In reality, molecules must satisfy multiple constraints simultaneously, including stability, selectivity, pharmacokinetics, toxicity, and synthetic feasibility.
When pipelines focus too heavily on a single metric, they often produce candidates that perform well computationally but fail in downstream validation.
This is one of the key reasons why high-scoring molecules frequently do not translate into viable drugs.
The Gap Between Prediction and Physics
Many AI models in drug discovery are fundamentally statistical. They learn patterns from data, but they do not inherently understand the physical principles governing molecular interactions.
As a result, predictions may overlook critical factors such as:
- protein flexibility
- solvent effects
- entropic contributions
- time-dependent interaction stability
These factors are central to real biological systems.
Without incorporating physics-based validation, AI-generated predictions can appear convincing but fail under realistic conditions. A molecule that seems to bind well in a static model may quickly lose stability when evaluated dynamically.
This gap between statistical prediction and physical reality is a major source of failure in AI-driven pipelines.
The Absence of Workflow Integration
Beyond individual model limitations, one of the most critical issues is the lack of integrated workflows.
Drug discovery is inherently iterative. Insights from one stage must inform decisions at the next, and feedback from experimental validation should continuously refine computational models.
In many AI pipelines, this feedback loop is either weak or entirely absent.
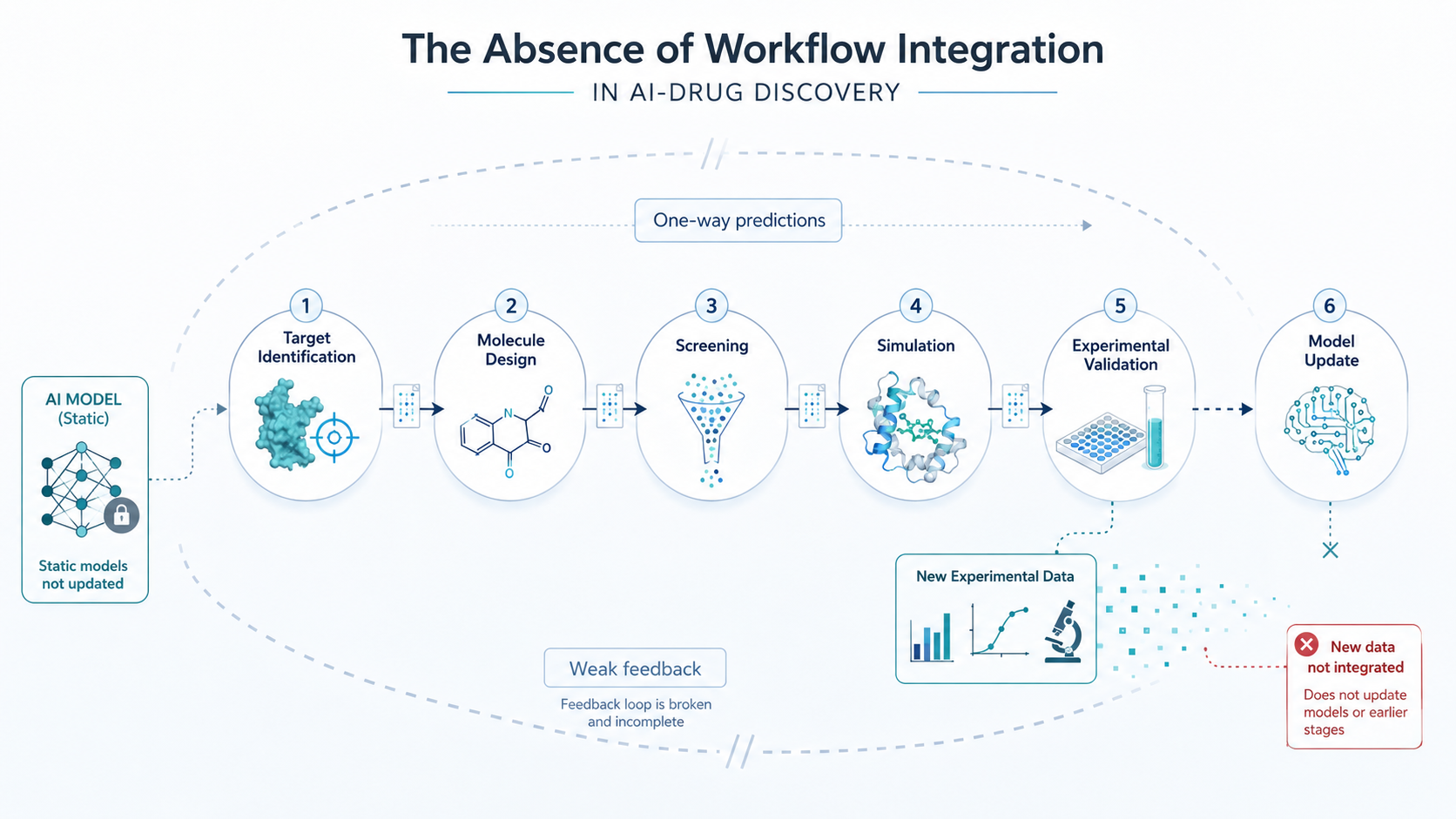
Predictions are generated, evaluated in isolation, and passed downstream without systematic integration. This prevents the pipeline from learning effectively and adapting to new information.
Without closed-loop workflows, AI systems remain static, even as new data becomes available.
The Data Problem: Quantity vs Quality
AI models are often described as data-driven, but not all data contributes equally to meaningful predictions.
Many pipelines rely on large, heterogeneous datasets that are:
- noisy
- incomplete
- biased toward specific targets or chemistries
In addition, much of the most valuable biological knowledge remains embedded in unstructured sources such as literature, patents, and experimental reports.
Without proper curation and contextualization, raw data can lead to misleading conclusions.
The challenge is not simply accessing data, but transforming it into structured, connected knowledge that can inform decision-making across the pipeline.
The Missing Link: Decision Intelligence
Ultimately, the goal of drug discovery is not prediction. It is decision-making.
Every stage of the pipeline involves choices:
- which targets to pursue
- which molecules to synthesize
- which candidates to advance
- which hypotheses to test
AI pipelines that focus solely on prediction often fail because they do not address this broader context.
What is needed is not just predictive intelligence, but decision intelligence—the ability to integrate multiple sources of information, evaluate trade-offs, and guide actions in a consistent and reliable manner.
Medvolt’s Approach: Building Integrated Discovery Pipelines
At Medvolt, the limitations of traditional AI pipelines are addressed through a fundamentally different design philosophy.
Rather than treating AI models as standalone tools, Medvolt’s platform is built as an integrated discovery system that connects multiple computational layers into a unified workflow.
This begins with knowledge-driven discovery, where biological insights are derived from curated datasets, knowledge graphs, multi-omics integration, and literature analysis. This ensures that target identification and hypothesis generation are grounded in connected biological context.
Molecule design is approached through AI-guided generation, but within constraints informed by structural biology, chemical feasibility, and downstream requirements. Generated molecules are not evaluated in isolation, but as part of a broader optimization process.
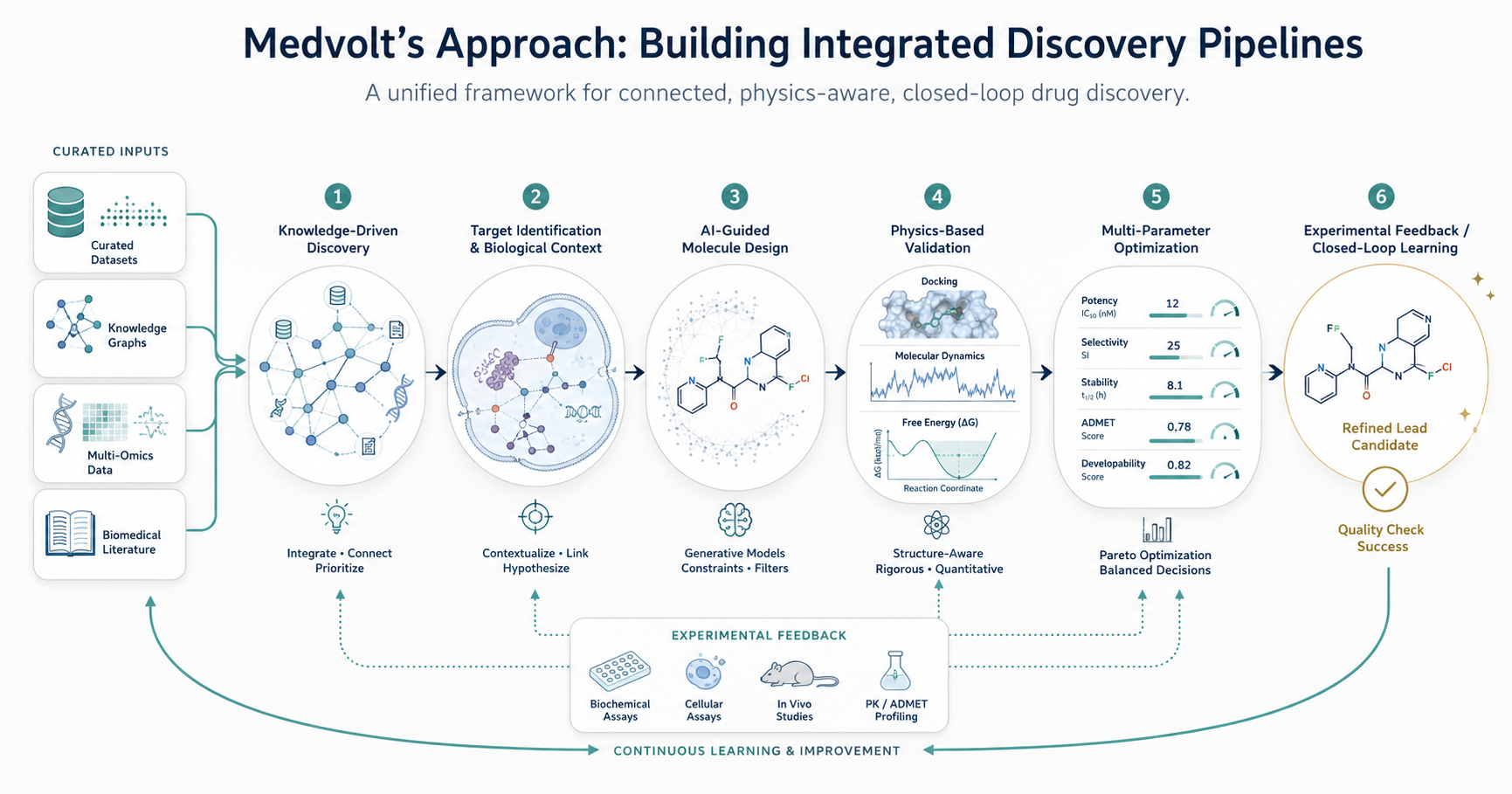
Crucially, predictions are continuously validated through physics-based methods, including docking, molecular dynamics simulations, and free energy calculations. This bridges the gap between statistical models and real molecular behavior.
The workflow is further strengthened by multi-parameter optimization, where candidates are evaluated across potency, selectivity, stability, and developability simultaneously, rather than sequentially.
Finally, Medvolt’s pipelines are designed as closed-loop systems, where computational predictions and experimental data inform each other iteratively. This allows the system to improve over time, rather than remaining static.
The result is not just a collection of AI models, but a cohesive framework for scientific decision-making.
Why This Matters for the Industry
As AI continues to evolve, the competitive landscape in drug discovery is shifting.
Access to models and data is becoming increasingly widespread. The true differentiator is no longer the presence of AI, but how effectively it is integrated into the discovery process.
Pipelines that remain fragmented, metric-driven, and disconnected from physical validation will continue to struggle with reproducibility and real-world impact.
In contrast, integrated systems that combine biological understanding, computational modeling, and iterative learning will be better positioned to deliver meaningful outcomes.
Conclusion
The failure of many AI drug discovery pipelines is not a reflection of the limitations of artificial intelligence itself.
It is a reflection of how these systems are designed and deployed.
Fragmentation, overreliance on narrow metrics, lack of physics-based validation, poor data integration, and absence of closed-loop workflows all contribute to the gap between promise and reality.
At Medvolt, this shift defines how we approach drug discovery. Because the future of computational pharma will not be built on models alone. It will be built on systems that can turn those models into reliable scientific outcomes.