Why the Future of Drug Discovery Depends on Integrated Systems, Not Isolated Intelligence
Drug discovery is shifting from isolated AI models to integrated systems that connect biology, chemistry, and simulations into decision-ready workflows.
Introduction
Artificial intelligence has rapidly become one of the defining technologies in modern drug discovery. Across the pharmaceutical and biotechnology industries, AI models are now used to predict protein structures, generate novel molecules, identify disease-associated targets, analyze biomedical literature, and optimize chemical properties at scales that were unimaginable a decade ago. The pace of progress has been extraordinary, and in many ways, AI has fundamentally reshaped what computational discovery can achieve.
Yet despite this momentum, a critical gap remains between prediction and practical impact.
Many organizations today possess powerful AI models, access to massive datasets, and significant computational infrastructure. However, only a small fraction are able to consistently translate those capabilities into reliable, scalable, and decision-ready drug discovery workflows. As a result, the industry is beginning to realize something important:
AI models alone are not enough. The core challenge in modern drug discovery is no longer simply building intelligent models. It is building systems that can connect those models into coherent scientific workflows capable of supporting real-world discovery programs from hypothesis generation to candidate optimization.
The Rise of AI-Centric Drug Discovery
Over the last decade, computational drug discovery has evolved from traditional cheminformatics and rule-based methods into a highly AI-driven ecosystem. Deep learning models can now infer protein structures with remarkable accuracy, generative systems can create entirely new molecular scaffolds, and large biological datasets can be mined for patterns that would be nearly impossible to identify manually.
This transformation has been accelerated by advances in high-performance computing, the growth of publicly available biomedical data, and improvements in machine learning architectures specifically tailored for chemistry and biology. As a result, AI has become central to the future vision of pharmaceutical R&D.
However, an important misconception often accompanies this progress. There is a growing tendency to assume that better models automatically translate into better discovery outcomes. In practice, this assumption rarely holds true.
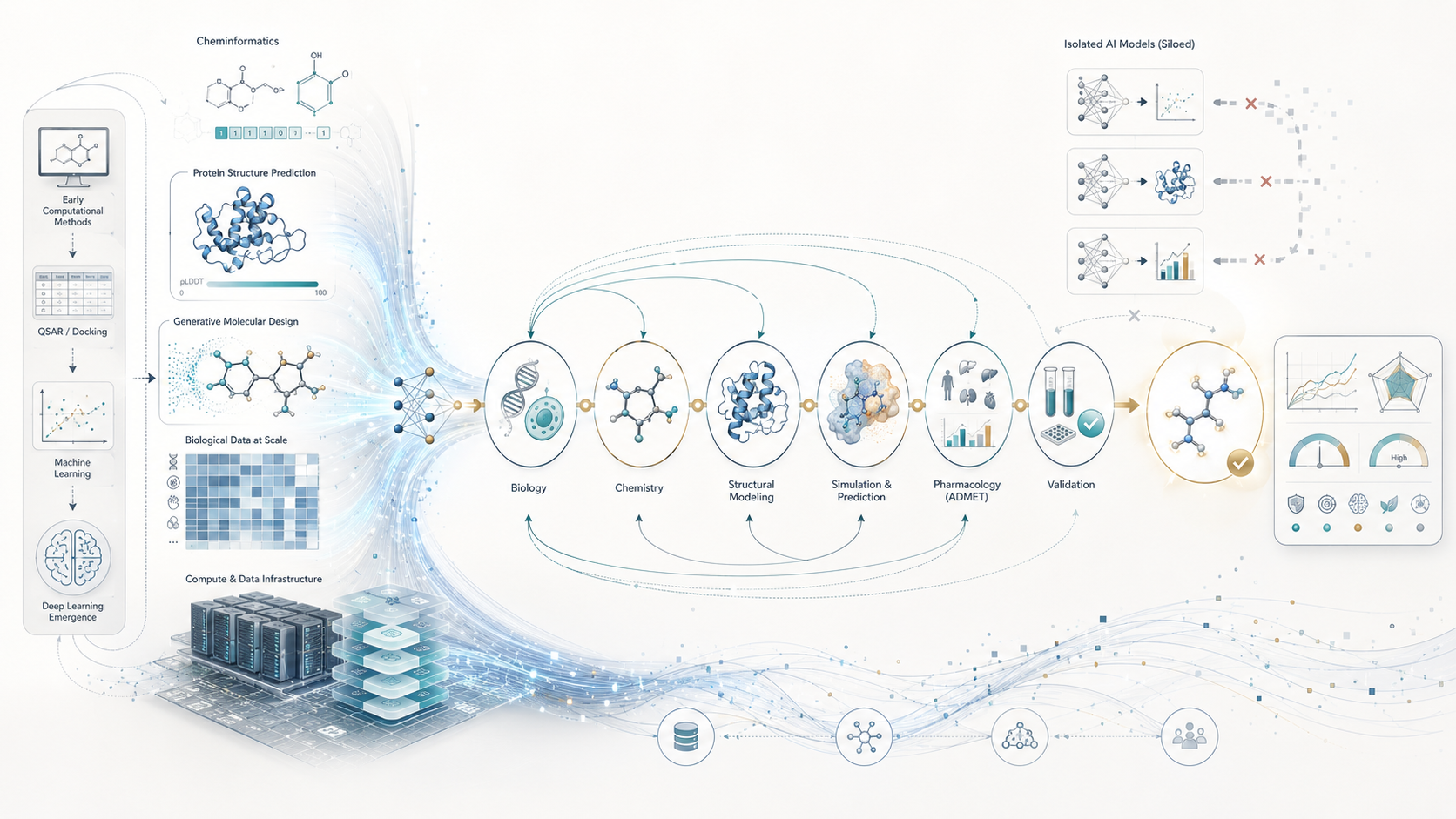
Drug discovery is not a single prediction problem. It is a complex sequence of interconnected scientific decisions spanning biology, chemistry, structural modeling, pharmacology, and translational validation. A molecule that appears promising in one computational context may completely fail when evaluated under another layer of the pipeline.
This is where many AI-driven systems begin to break down.
The Problem with Isolated Intelligence
One of the most common weaknesses in computational drug discovery today is fragmentation. Models are often developed independently, optimized for narrow objectives, and deployed as isolated tools rather than integrated systems.
A generative AI model may design a chemically novel molecule with strong predicted affinity toward a target protein. Yet that same molecule may:
- exhibit poor ADMET properties
- fail molecular dynamics stability analysis
- become synthetically impractical
- lack selectivity across related targets
- perform poorly under realistic biological conditions
In many organizations, these problems are compounded by disconnected workflows. Outputs from one computational stage are manually transferred into another system, interpreted independently, and validated using inconsistent assumptions. As the complexity of discovery programs increases, these disconnected processes become increasingly difficult to scale.
This creates a paradox that is becoming more visible across the industry: organizations possess more computational capability than ever before, yet meaningful improvements in R&D productivity remain difficult to achieve.
The issue is not necessarily the quality of individual models. The issue is the absence of integrated scientific workflows that can connect predictions into a coherent decision-making process.
Drug Discovery Is a Workflow Problem
The central challenge in modern computational discovery is not simply prediction accuracy. It is workflow orchestration.
Effective drug discovery requires the ability to connect multiple layers of reasoning into a unified process. Biological context, structural understanding, molecular design, simulation, optimization, and validation all need to interact continuously rather than operate as isolated stages.
This distinction is extremely important.
An isolated AI model may answer a narrow question effectively. A workflow determines whether that answer is actionable within the broader scientific context.
For example, discovering a molecule with strong predicted affinity is useful only if the same molecule also demonstrates acceptable stability, selectivity, developability, and pharmacological behavior. Similarly, identifying a promising target is valuable only if that insight can be connected to realistic therapeutic strategies and experimentally viable pathways.
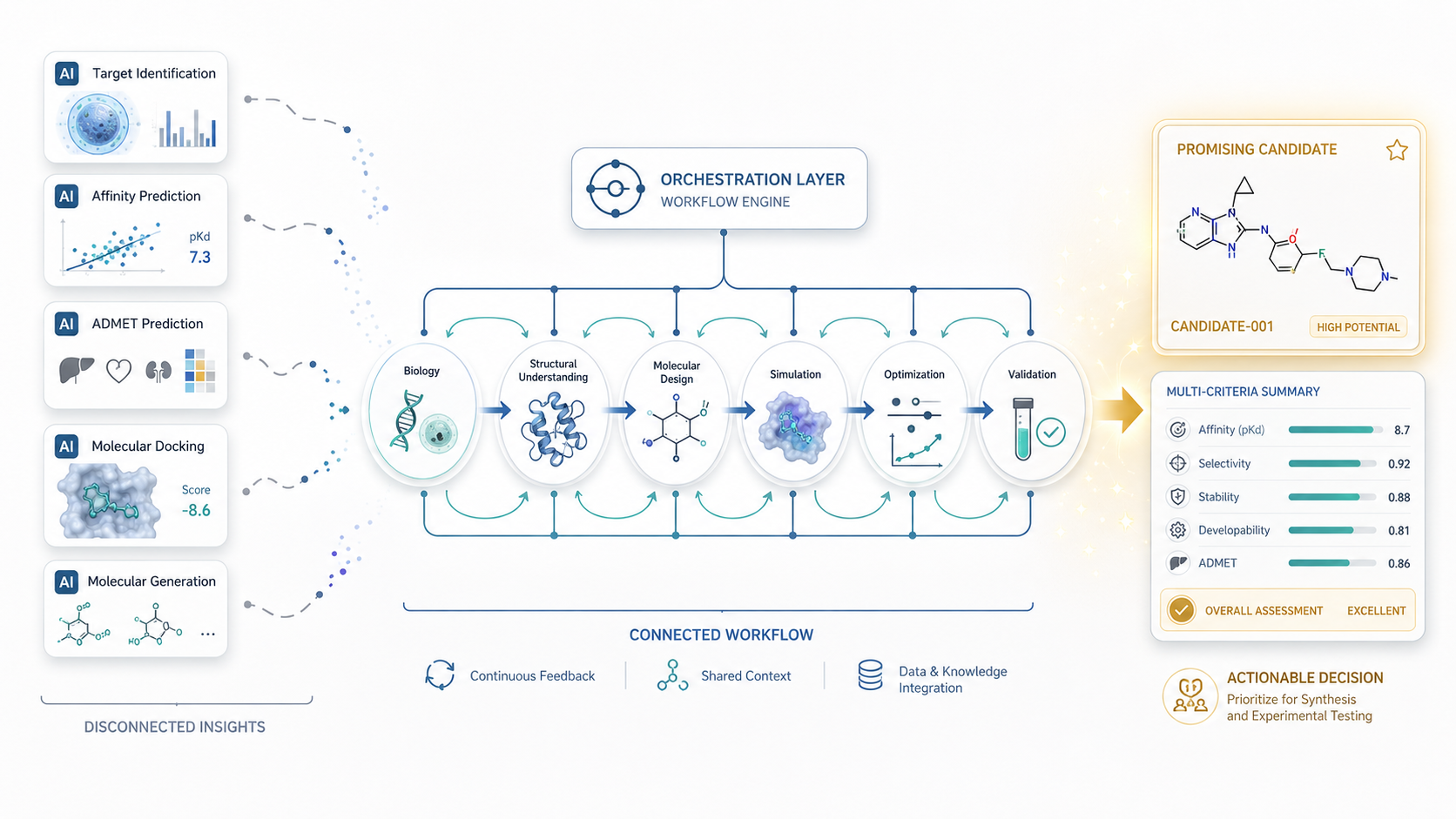
Without integrated workflows, computational predictions often become disconnected insights rather than usable scientific decisions.
Why Integrated Workflows Matter
Integrated workflows solve several of the most persistent challenges in computational drug discovery.
First, they preserve context across discovery stages. Biological relationships identified during target discovery can continue informing downstream molecular design and optimization rather than being lost between disconnected systems.
Second, integrated workflows improve consistency. When multiple computational stages operate within a unified framework, predictions can be evaluated under coherent scientific assumptions rather than fragmented scoring systems and incompatible methodologies.
Third, workflows allow for true multi-parameter optimization. Drug candidates rarely fail because of a single issue. They fail because multiple biological, chemical, and pharmacological constraints interact simultaneously. Integrated systems allow these competing factors to be evaluated together rather than sequentially.
Most importantly, workflows enable continuous learning. Modern drug discovery increasingly depends on closed-loop systems where computational predictions inform experiments, and experimental feedback subsequently improves future predictions. Without integration, this learning cycle becomes inefficient and difficult to scale.
The Industry Is Moving Toward Systems-Level Discovery
The pharmaceutical industry is gradually shifting away from tool-centric thinking toward systems-level discovery architectures. This transition reflects a broader understanding that biological complexity cannot be addressed through isolated optimizations alone.
Modern discovery programs increasingly involve multi-dimensional scientific challenges, such as:
- precision oncology
- synthetic lethality
- protein engineering
- multi-target therapeutics
- biologics and enzyme therapeutics
These areas require workflows capable of connecting biology, chemistry, structural modeling, and simulation in a coordinated manner.
At the same time, many AI capabilities are becoming commoditized. Public foundation models, open-source frameworks, and large shared datasets are reducing barriers to entry across the industry. In this environment, competitive advantage is shifting away from possessing individual AI models toward building systems capable of orchestrating them effectively.
The future will likely belong to organizations that can transform fragmented computational capabilities into unified scientific intelligence.
Medvolt’s Approach: From Models to Connected Discovery Systems
At Medvolt, this systems-level philosophy is central to how computational discovery workflows are designed.
Rather than treating AI as a standalone capability, Medvolt’s platform architecture was developed to connect multiple computational layers into an integrated discovery ecosystem. The goal is not simply to generate predictions, but to improve scientific decision-making throughout the entire pipeline.
This begins with knowledge-driven discovery. Biological hypotheses are contextualized through integration of biomedical literature, knowledge graphs, pathway relationships, multi-omics data, and disease-specific insights. Instead of treating data as isolated information sources, the platform connects genes, proteins, compounds, pathways, and diseases into a unified framework that supports deeper biological reasoning.
Molecule design workflows are similarly integrated. Generative AI models operate within structural, chemical, and developability constraints so that generated compounds remain biologically meaningful and experimentally realistic. These molecules are not evaluated purely through static scoring functions. They progress through structure-aware workflows involving docking, molecular dynamics simulations, and free energy calculations to assess interaction stability and thermodynamic feasibility under realistic conditions.
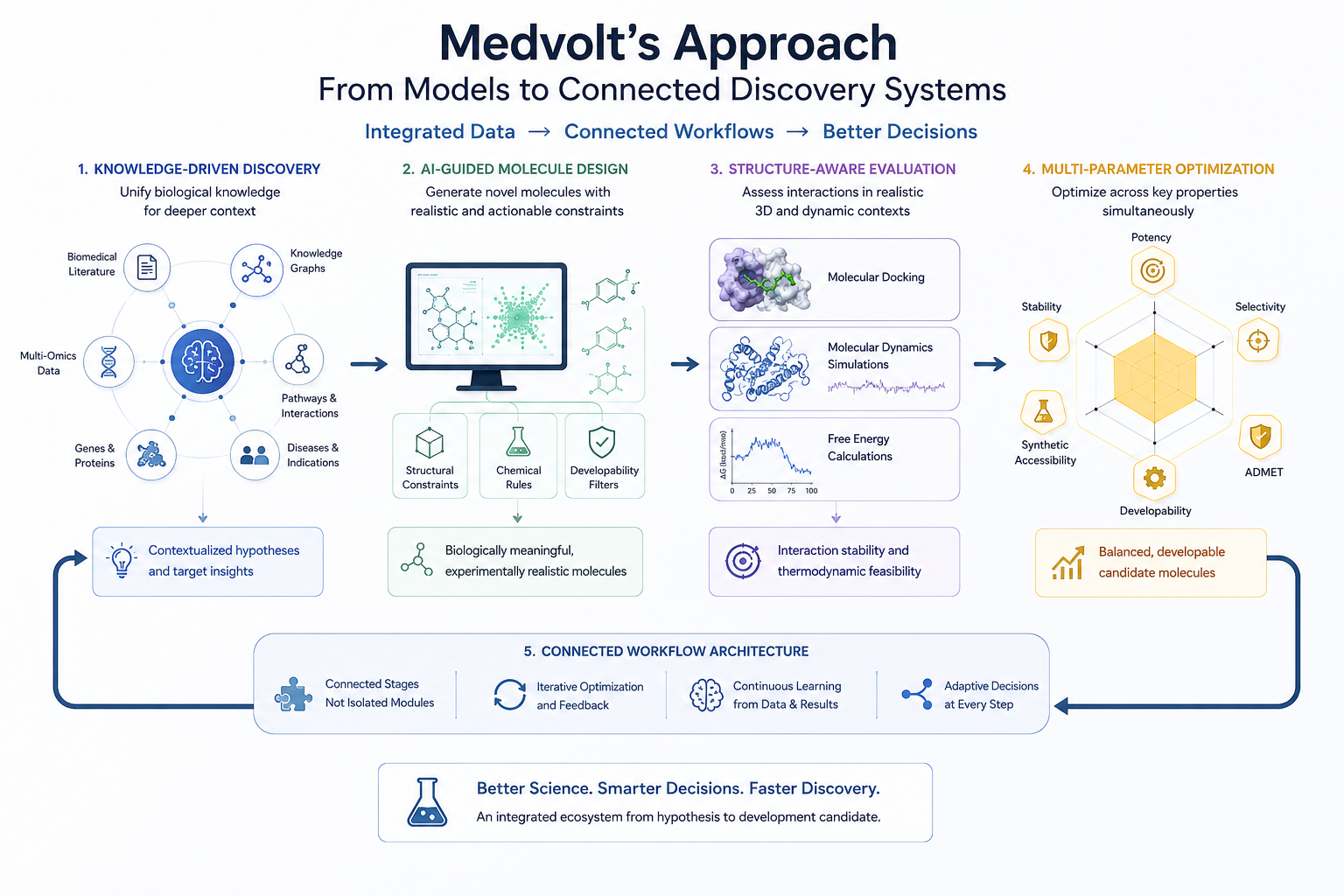
Importantly, these computational layers are not isolated modules operating independently. They function as connected stages within a larger workflow architecture designed to support iterative optimization and continuous refinement.
This integration becomes especially important in complex therapeutic areas where biological uncertainty, dynamic molecular behavior, and multi-parameter optimization all intersect. In such environments, isolated predictions have limited value unless they can be interpreted within a broader systems-level context.
Why Workflow Architecture Will Define the Future
As AI adoption accelerates across life sciences, the differentiator will increasingly shift away from access to models and toward the ability to orchestrate workflows effectively.
The future of computational drug discovery will depend on systems capable of integrating multiple scientific dimensions simultaneously while maintaining continuity across the discovery lifecycle. Organizations that succeed will likely be those that can combine predictive modeling, structural biology, physics-based simulations, biological reasoning, and experimental feedback into unified and scalable architectures.
This transition represents a broader evolution in how discovery itself is understood. The industry is moving away from isolated computational predictions toward connected scientific intelligence.
In this new paradigm, workflows are no longer operational details. They are the foundation of effective discovery.
Conclusion
Artificial intelligence is transforming drug discovery, but models alone are not enough.
Drug discovery is fundamentally a workflow-driven challenge involving interconnected scientific decisions across biology, chemistry, structural modeling, simulation, and translational validation. Without integration, even highly advanced AI models remain limited in practical impact.
The next generation of discovery platforms will not be defined solely by prediction accuracy. They will be defined by how effectively they connect multiple layers of scientific reasoning into coherent systems capable of supporting real-world decision-making.
At Medvolt, this belief shapes the foundation of our computational workflows. Because in modern drug discovery, the true challenge is not simply generating predictions.
It is transforming those predictions into reliable scientific outcomes.
And that requires workflows, not just models.